Thomas Gingeras did not intend to upend basic ideas about how the human body works. In 2012 the geneticist, now at Cold Spring Harbor Laboratory in New York State, was one of a few hundred colleagues who were simply trying to put together a compendium of human DNA functions. Their project was called ENCODE, for the Encyclopedia of DNA Elements. About a decade earlier almost all of the three billion DNA building blocks that make up the human genome had been identified. Gingeras and the other ENCODE scientists were trying to figure out what all that DNA did.
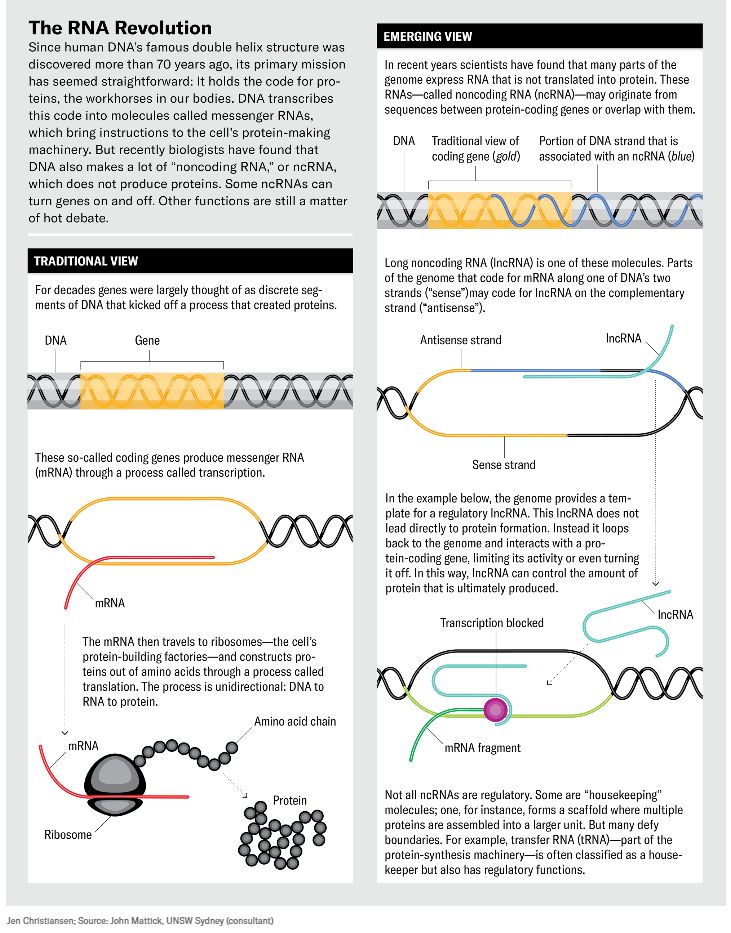
The assumption made by most biologists at that time was that most of it didn’t do much. The early genome mappers estimated that perhaps 1 to 2 percent of our DNA consisted of genes as classically defined: stretches of the genome that coded for proteins, the workhorses of the human body that carry oxygen to different organs, build heart muscles and brain cells, and do just about everything else people need to stay alive. Making proteins was thought to be the genome’s primary job. Genes do this by putting manufacturing instructions into messenger molecules called mRNAs, which in turn travel to a cell’s protein-making machinery. As for the rest of the genome’s DNA? The “protein-coding regions,” Gingeras says, were supposedly “surrounded by oceans of biologically functionless sequences.” In other words, it was mostly junk DNA.
So it came as rather a shock when, in several 2012 papers in Nature, he and the rest of the ENCODE team reported that at one time or another, at least 75 percent of the genome gets transcribed into RNAs. The ENCODE work, using techniques that could map RNA activity happening along genome sections, had begun in 2003 and came up with preliminary results in 2007. But not until five years later did the extent of all this transcription become clear. If only 1 to 2 percent of this RNA was encoding proteins, what was the rest for? Some of it, scientists knew, carried out crucial tasks such as turning genes on or off; a lot of the other functions had yet to be pinned down. Still, no one had imagined that three quarters of our DNA turns into RNA, let alone that so much of it could do anything useful.
Some biologists greeted this announcement with skepticism bordering on outrage. The ENCODE team was accused of hyping its findings; some critics argued that most of this RNA was made accidentally because the RNA-making enzyme that travels along the genome is rather indiscriminate about which bits of DNA it reads.
Now it looks like ENCODE was basically right. Dozens of other research groups, scoping out activity along the human genome, also have found that much of our DNA is churning out “noncoding” RNA. It doesn’t encode proteins, as mRNA does, but engages with other molecules to conduct some biochemical task. By 2020 the ENCODE project said it had identified around 37,600 noncoding genes—that is, DNA stretches with instructions for RNA molecules that do not code for proteins. That is almost twice as many as there are protein-coding genes. Other tallies vary widely, from around 18,000 to close to 96,000. There are still doubters, but there are also enthusiastic biologists such as Jeanne Lawrence and Lisa Hall of the University of Massachusetts Chan Medical School. In a 2024 commentary for the journal Science, the duo described these findings as part of an “RNA revolution.”
What makes these discoveries revolutionary is what all this noncoding RNA—abbreviated as ncRNA—does. Much of it indeed seems involved in gene regulation: not simply turning them off or on but also fine-tuning their activity. So although some genes hold the blueprint for proteins, ncRNA can control the activity of those genes and thus ultimately determine whether their proteins are made. This is a far cry from the basic narrative of biology that has held sway since the discovery of the DNA double helix some 70 years ago, which was all about DNA leading to proteins. “It appears that we may have fundamentally misunderstood the nature of genetic programming,” wrote molecular biologists Kevin Morris of Queensland University of Technology and John Mattick of the University of New South Wales in Australia in a 2014 article.
Another important discovery is that some ncRNAs appear to play a role in disease, for example, by regulating the cell processes involved in some forms of cancer. So researchers are investigating whether it is possible to develop drugs that target such ncRNAs or, conversely, to use ncRNAs themselves as drugs. If a gene codes for a protein that helps a cancer cell grow, for example, an ncRNA that shuts down the gene might help treat the cancer.
A few noncoding RNAs had been known for many decades, but those seemed to have some role in protein manufacture. For instance, only a few years after Francis Crick, James Watson and several of their colleagues deduced the structure of DNA, researchers found that some RNA, called transfer RNA, grabs onto amino acids that eventually get strung together into proteins.
In the 1990s, however, scientists realized ncRNA could do things quite unrelated to protein construction. These new roles came to light from efforts to understand the process of X-inactivation, wherein one of the two X chromosomes carried by females is silenced, all 1,000 or so of its genes (in humans) being turned off. This process seemed to be controlled by a gene called XIST. But attempts to find the corresponding XIST protein consistently failed.
The reason, it turned out, was that the gene did not work through a protein but instead did so by producing a long noncoding (lnc) RNA molecule. Such RNAs are typically longer than about 200 nucleotides, which are the chemical building blocks of DNA and RNA. Using a microscopy technique called fluorescence in situ hybridization, Lawrence and her colleagues showed that this RNA wraps itself around one X chromosome (selected at random in each cell) to induce persistent changes that silence the genes. “This was the first evidence of a lncRNA that does something,” Lawrence says, “and it was totally surprising.”
If noncoding RNAs power the way a cell processes genetic information, it is possible they can be used in medicine.
XIST isn’t that unusual in generating an ncRNA, though. In the early 2000s it became clear that transcription of noncoding DNA sequences is widespread. For example, in 2002 a team at biotech company Affymetrix in Santa Clara, Calif., led by Gingeras, who was working there at the time, reported that much more of human chromosomes 21 and 22 gets transcribed than just the protein-coding regions.
It was only after ENCODE published its results in 2012, however, that ncRNA became impossible to ignore. Part of the antipathy toward those findings, says Peter Stadler, a bioinformatics expert at Leipzig University in Germany, is that they seemed like an unwanted and unneeded complication. “The biological community figured we already knew how the cell works, and so the discovery of was more of an annoyance,” he says. What’s more, it showed that simpler organisms were not always a reliable guide to human biology: there is far less ncRNA in bacteria, studies of which had long shaped thinking about how genes are regulated.
But now there is no turning back the tide: many thousands of human lncRNAs have been reported, and Mattick suspects the real number is greater than 500,000. Yet only a few of these have been shown to have specific functions, and how many of them really do remains an open question. “I personally don’t think all of those RNAs have an individual role,” Lawrence says. Some, though, may act in groups to regulate other molecules.
How lncRNAs perform such regulation is also still a matter of debate. One idea is that they help to form so-called condensates: dense fluid blobs containing a range of different regulatory molecules. Condensates are thought to hold all the relevant players in one place long enough for them to do their job collectively. Another idea is that lncRNAs affect the structure of chromatin—the combination of DNA and proteins that makes up chromosome fibers in the cell nucleus. How chromatin is structured determines which of its genes are accessible and can be transcribed; if parts of chromatin are too tightly packed, the enzyme machinery of transcription can’t reach it. “Some lncRNAs appear to be involved with chromatin-modifying complexes,” says Marcel Dinger, a genomics researcher at the University of Sydney.
If only 1 to 2 percent of the RNA from our genome was encoding proteins, what was the rest for? Some, scientists knew, carried out crucial tasks such as turning genes on or off.
Lawrence and Hall suspect that lncRNAs could supply scaffolds for organizing other molecules, for example, by holding some of the many hundreds of RNA-binding proteins in functional assemblies. One lncRNA called NEAT1, which is involved in the formation of small compartments in the nucleus called paraspeckles, has been shown capable of binding up to 60 of these proteins. Or such RNA scaffolding could arrange chromatin itself into particular structures and thereby affect gene regulation. Such RNA scaffolding could have regularly repeating modules and thus repetitive sequences—a feature that has long been regarded as a hallmark of junk DNA but lately is appearing to be not so junky after all. This view of lncRNA as scaffolding is supported by a 2024 report of repeat-rich ncRNAs in mouse brain cells that persist for at least two years. The research, by Sara Zocher of the German Center for Neurodegenerative Diseases in Dresden and her co-workers. found these ncRNAs seem to be needed to keep parts of chromatin in a compact and silent state.
These lncRNAs are just one branch of the noncoding RNA family, and biologists keep discovering others that appear to have different functions and different ways of affecting what happens to a cell—and thus the entire human body.
Some of these RNAs are not long at all but surprisingly short. Their story began in the 1980s, when Victor Ambros, working as a postdoctoral researcher in the laboratory of biologist Robert Horvitz at the Massachusetts Institute of Technology, was studying a gene denoted lin-4 in the worm Caenorhabditis elegans. Mutations of lin-4 caused developmental defects in which “the cells repeated whole developmental programs that they should have transitioned beyond,” says Ambros, now at the University of Massachusetts Medical School. It seemed that lin-4 might be a kind of “master regulator” controlling the timing of different stages of development.
“We thought lin-4 would be a protein-coding gene,” Ambros says. To figure out what role this putative protein plays, Ambros and his colleagues cloned the C. elegans gene and looked at its product—and found that the effects of the gene may not be mediated by any protein but by the gene’s RNA product alone. This molecule looked ridiculously short: just 22 nucleotides long, a mere scrap of a molecule for such big developmental effects.
This was the first known microRNA (miRNA). At first “we thought this might be a peculiar characteristic of C. elegans,” Ambros says. But in 2000 Gary Ruvkun, another former postdoc in the Horvitz lab, and his co-workers found that another of these miRNA genes in C. elegans, called let-7, appears in essentially identical form in many other organisms, including vertebrates, mollusks and insects. This implies that it is a very ancient gene and “must have been around for 600 million to 700 million years” before these diverse lineages went their separate ways, Ambros says. If miRNAs are so ancient, “there had to be others out there.”
Indeed, there are. Today more than 2,000 miRNAs have been identified in the human genome, generally with regulatory roles. One of the main ways miRNAs work is by interfering with the translation of a gene’s mRNA transcript into its corresponding protein. Typically the miRNA comes from a longer molecule, perhaps around 70 nucleotides long, known as pre-miRNA. This molecule is seized by an enzyme called Dicer, which chops it into smaller fragments. These pieces, now miRNAs, move to a class of proteins called Argonautes, components of a protein assembly called the RNA-induced silencing complex (RISC). The miRNAs guide the RISC to an mRNA, and this either stops the mRNA from being translated into a protein or leads to its degradation, which has the same effect. This regulatory action of miRNAs guides processes ranging from the determination of cell “fate” (the specialized cell types they become) to cell death and management of the cell cycle.
Key insights into how such small RNAs can regulate other RNA emerged from studies in C. elegans in 1998 by molecular biologists Andrew Fire, Craig Mello and their co-workers, for which Fire and Mello were awarded the 2006 Nobel Prize in Physiology or Medicine. They learned that RISC is guided by slightly different RNA strands named small interfering (si) RNA. The process ends with the mRNA being snipped in half, a process called RNA interference.
MiRNAs do pose a puzzle, however. A given miRNA typically has a sequence that matches up with lots of mRNAs. How, then, is there any selectivity about which genes they silence? One possibility is that miRNAs work in gangs, with several miRNAs joining forces to regulate a given gene. The different combinations, rather than individual snippets, are what match specific genes and their miRNAs.
Why would miRNA gene regulation work in this complicated way? Ambros suspects it might allow for “evolutionary fluidity”: the many ways in which different miRNAs can work together, and the number of possible targets each of them can have, offer a lot of flexibility in how genes are regulated and thus in what traits might result. That gives an organism many evolutionary options, so that it is more able to adapt to changing circumstances.
One class of small RNAs regulates gene expression by directly interfering with transcription in the cell nucleus, triggering mRNA degradation. These PIWI-interacting (pi) RNAs work in conjunction with a class of proteins called PIWI Argonautes. PiRNAs operate in germline cells (gametes), where they combat “selfish” DNA sequences called transposons or “jumping genes”: sequences that can insert copies of themselves throughout the genome in a disruptive way. Thus, piRNAs are “a part of the genome’s immune system,” says Julius Brennecke of the Institute of Molecular Biotechnology of the Austrian Academy of Sciences. If the piRNA system is artificially shut down, “the gametes’ genomes are completely shredded, and the organism is completely sterile,” he says.
Still other types of ncRNAs, called small nucleolar RNAs, work within cell compartments called nucleoli to help modify the RNA in ribosomes—a cell’s protein-making factories—as well as transfer RNA and mRNA. These are all ways to regulate gene expression. Then there are circular RNAs: mRNA molecules (particularly in neurons) that get stitched into a circular form before they are moved beyond the nucleus into the cytoplasm. It’s not clear how many circular RNAs are important—some might just be transcriptional “noise”—but there is some evidence that at least some of them have regulatory functions.
In addition, there are vault RNAs that help to transport other molecules within and between cells, “small Cajal-body-specific RNAs” that modify other ncRNAs involved in RNA processing, and more. The proliferation of ncRNA varieties lends strength to Mattick’s claim that RNA, not DNA, is “the computational engine of the cell.”
If ncRNAs indeed power the way a cell processes genetic information, it is possible they can be used in medicine. Disease is often the result of a cell doing the wrong thing because it gets the wrong regulatory instructions: cells that lose proper control of their cycle of growth and division can become tumors, for example. Currently medical efforts to target ncRNAs and alter their regulatory effects often use RNA strings called antisense oligonucleotides (ASOs). These strands of nucleic acid have sequences that are complementary to the target RNA, so they will pair up with and disable it. ASOs have been around since the late 1970s. But it has been hard to make them clinically useful because they get degraded quickly in cells and have a tendency to bind to the wrong targets, with potentially drastic consequences.
Some ASOs, however, are being developed to disable lncRNAs that are associated with cancers such as lung cancer and acute myeloid leukemia. Other lncRNAs might act as drugs themselves. One known as MEG3 has been found, preliminarily, to act as a tumor suppressor. Small synthetic molecules, which are easier than ASOs to fine-tune and deliver into the body as pharmaceuticals, are also being explored for binding to lncRNAs or otherwise inhibiting their interactions with proteins. Getting these approaches to work, however, has not been easy. “As far as I am aware, no lncRNA target or therapeutic has entered clinical development,” Gingeras says.

James Yang
Targeting the smaller regulatory RNAs such as miRNAs might prove more clinically amenable. Because miRNAs typically hit many targets, they can do many things at once. For example, miRNAs in families denoted miR-15a and miR-16-1 act as tumor suppressors by targeting several genes that themselves suppress cell death (apoptosis, a defense against cancer) and are being explored for cancer therapies.
Yet a problem with using small RNAs as drugs is that they elicit an immune response. Precisely because the immune system aims to protect against viral RNA, it usually recognizes and attacks any “nonself” RNA. One strategy for protecting therapeutic RNA from immune assault and degradation is to chemically modify its backbone so that it forms a nonnatural “locked” ring structure that the degrading enzymes can’t easily recognize.
Some short ASOs that target RNAs are already approved for clinical use, such as the drugs inotersen to treat amyloidosis and golodirsen for Duchenne muscular dystrophy. Researchers are also exploring antisense RNAs fewer than 21 nucleotides long that target natural regulatory miRNAs because it is only beyond that length that an RNA tends to trigger an immune reaction.
These are early days for RNA-based medicine, precisely because the significance of ncRNA itself in human biology is still relatively new and imperfectly understood. The more we appreciate its pervasive nature, the more we can expect to see RNA being used to control and improve our well-being. Nils Walter of the Center for RNA Biomedicine at the University of Michigan wrote in an article early in 2024 that the burgeoning promise of RNA therapeutics “only makes the need for deciphering ncRNA function more urgent.” Succeeding in this goal, he adds, “would finally fulfill the promise of the Human Genome Project.”
Despite this potential of noncoding RNA in medicine, the debate continues about how much of it truly matters for our cells. Geneticists Chris Ponting of the University of Edinburgh and Wilfried Haerty of the Earlham Institute in Norwich, England, are among the skeptics. In a 2022 article they argued that most lncRNAs are just “transcriptional noise,” accidentally transcribed from random bits of DNA. “Relatively few human lncRNAs ... contribute centrally to human development, physiology, or behavior,” they wrote.
Brennecke advises caution about current high estimates of the number of noncoding genes. Although he agrees that such genes “have been underappreciated for a long time,” he says we should not leap to assuming that all lncRNAs have functions. Many of them are transcribed only at low levels, which is what one would expect if indeed they were just random noise. Geneticist Adrian Bird of the University of Edinburgh points out that the abundance of the vast majority of ncRNAs seems to be well below one molecule per cell. “It is difficult to see how essential functions can be exerted by an ncRNA if it is absent in most cells,” he says.
But Gingeras counters that this low expression rate might reflect the very tissue-specific roles of ncRNAs. Some, he says, are expressed more in one part of a tissue than in another, suggesting that expression levels in each cell are sensitive to signals coming from surrounding tissues. Lawrence points out that, despite the low expression levels, there are often shared patterns of expression across cells of a particular type, making it harder to argue that the transcription is simply random. And Hall doubts that cells are really so prone to “bad housekeeping” that they will habitually churn out lots of useless RNA. Lawrence and Hall’s suggestion that some lncRNAs have collective effects on chromatin structure would mean that no individual one of them is needed at high expression levels and that their precise sequence doesn’t matter too much.
That lack of specificity in sequence and binding targets, Dinger says, means that a mutation of a nucleotide in an ncRNA typically won’t have the same negative impact on its function as it tends to in a protein-coding DNA sequence. So it would not be surprising to see quite a lot of sequence variation. Dinger argues that it makes more sense to assume that “genetically encoded molecules are potentially functional until shown otherwise, rather than junk unless proven functional.” Some in the ENCODE team now agree that not all of the 75 percent or so of human genome transcription might be functionally significant. But many researchers make the point that surely many more of the noncoding molecules do meaningful things than was suspected before.
Demonstrating functional roles for lncRNAs is often tricky. In part, Gingeras says, this may be because lncRNA might not be the biochemically active molecule in a given process: it might be snipped up into short RNAs that actually do the work. But because long and short RNAs tend to be characterized via different techniques, researchers may end up searching for the wrong thing. What’s more, long RNAs are often cut up into fragments and then spliced back together again in various combinations, the exact order often depending on the condition of the host cell.
At its roots, the controversy over noncoding RNA is partly about what qualifies a molecule as “functional.” Should the criterion be based on whether the sequence is maintained between different species? Or whether deleting the molecule from an organism’s repertoire leads to some observable change in a trait? Or simply whether it can be shown to be involved in some biochemical process in the cell? If repetitive RNA acts collectively as a chromosome “scaffold” or if miRNAs act in a kind of regulatory swarm, can any individual one of them really be considered to have a “function”?
Gingeras says he is perplexed by ongoing claims that ncRNAs are merely noise or junk, as evidence is mounting that they do many things. “It is puzzling why there is such an effort to persuade colleagues to move from a sense of interest and curiosity in the ncRNA field to a more dubious and critical one,” he says.
Perhaps the arguments are so intense because they undercut the way we think our biology works. Ever since the epochal discovery about DNA’s double helix and how it encodes information, the bedrock idea of molecular biology has been that there are precisely encoded instructions that program specific molecules for particular tasks. But ncRNAs seem to point to a fuzzier, more collective, logic to life. It is a logic that is harder to discern and harder to understand. But if scientists can learn to live with the fuzziness, this view of life may turn out to be more complete.
Philip Ball is a science writer and former Nature editor based in London. His most recent book is How Life Works (University of Chicago Press, 2023).
Scientific American covers the most important and exciting research, ideas and knowledge in science, health, technology, the environment and society. It is committed to sharing trustworthy knowledge, enhancing our understanding of the world, and advancing social justice.
History
Founded 1845, Scientific American is the oldest continuously published magazine in the United States. It has published articles by more than 200 Nobel Prize winners.Reach
Scientific American reaches more than 10 million people around the world each month through its website, print and digital editions, newsletters and app and millions more through social media and other platforms.
Authors Scientific American publishes work by journalists, scientists, scholars, policy makers and people with lived experience of scientific or social issues. See our submission guidelines for more information.
Content Scientific American publishes in-depth feature articles, timely news stories, expert opinion and commentary, podcasts, videos, multimedia, photo features and other forms of storytelling. In addition to our monthly flagship magazine and pieces published daily on our website, we produce special Collector’s Editions, the digital magazines Mind, Space & Physics and Health & Medicine, ebooks and other products. We also host live events and webinars featuring a diversity of experts and journalists who share their knowledge about current events the way that science and scholarly inquiry enhance our understanding of the world and shape our lives.
Subscribe to Scientific American
Science Journalism You Can Trust
Since 1845 Scientific American has been bringing its readers unique insights about developments in science and technology.
Insight, Mystery, Discovery
Stay up to date with award-winning reports on the latest research, ideas and knowledge in science.
Risk Free, Cancel Anytime
Subscribe today for full access to all of our stories. Cancel at any time for any reason.

Spots, Stripes and More: Working out the Logic of Animal Patterns
By Amber Dance
Knowable Magazine
More than 70 years ago, mathematician Alan Turing proposed a mechanism that explained how patterns could emerge from bland uniformity. Scientists are still using his model — and adding new twists — to gain a deeper understanding of animal markings.
May 23, 2024

Spread the word